Enzyme Stability Prediction
The QuData team conducted a study on predicting the thermostability of enzymes — key biological catalysts with important applications in medicine, the food industry, and agriculture. To address this challenge, we developed an ensemble approach combining neural networks (3D-ResNet), structural descriptors from MoleculeKit, and Rosetta scores. This combination enables accurate prediction of protein stability based on amino acid sequences and structural features, improving understanding of enzyme behavior and enhancing their efficiency in industrial applications and protein engineering.
 Business Challenge
Business Challenge
Enzymes are proteins that serve as catalysts in the chemical reactions within living organisms. They play a fundamental role in various biological processes and have applications in many different industries. Enzymes are used in food and beverage production, improving the quality of products. They are used in the development of pharmaceuticals and are vital in converting biomass into biofuels. These are just a few instances of the diverse applications of enzymes.
However, it's important to note that many enzymes face challenges related to stability, particularly in harsh conditions. This instability not only impacts their performance but also limits the amount of protein that cells can produce. Therefore, there's a pressing need for efficient computational methods to predict and enhance protein stability, representing a significant area of interest in both technical and scientific realms.
Comprehending and accurately predicting protein stability is a fundamental problem in biotechnology. This knowledge has far-reaching applications, from enhancing enzyme performance to addressing global challenges like sustainability and carbon neutrality. Enhancing enzyme stability has the potential to reduce costs and accelerate the pace at which scientists can experiment with new ideas.
 Solution Overview
Solution Overview
The objective of the research was to forecast the thermostability of enzyme variants. The thermostability data, experimentally measured (melting temperature), encompasses both natural sequences and sequences that have been engineered with single or multiple mutations based on the natural sequences.
A successful solution to this task may address the fundamental challenge of enhancing protein stability, enabling the faster and cost-efficient design of novel proteins, including valuable enzymes and therapeutics.
During research our team initially used a specific approach called "Thermonet," which showed promising results.
We experimented with different neural network architectures and settled on a simplified one that yielded a good score. To further improve our predictions, we used "molekulekit" utility to create feature descriptors and enhanced the model's performance.
Additionally, the QuData team found thatOverall, this study emphasizes the significance of Rosetta scores and the average of the hydrogen bond donor parameter in predicting thermal stability of enzymes.
Our research was instigated by a competition initiated by Novozymes. Participants were asked to create a model for predicting and ranking the thermostability of enzyme variants based on experimental melting temperature data from Novozymes' high throughput screening lab. You can learn more about the competition and its results here.


 Technical Details
Technical Details
In the pursuit of advancing predictions for thermal stability, the QuData team explored various strategies and technologies. The following summarizes the key components of our approach.
Initial Approach – nesp-thermonet: we began by recognizing the significant predictive capability of the "nesp-thermonet" approach.
Exploration of neural network architectures: we delved into the details provided in this article, which spurred our exploration of different neural network architectures. Ultimately, we converged on the Resnet-3D model.
Network pruning and simplification: to further dissect the architecture and pinpoint the key contributors to the prediction, we adopted an iterative pruning process, leading to the development of a simplified architecture. This architecture has the same input dimensions as the "Thermonet" work (14,16,16,16).
Leveraging feature descriptors: to enhance our predictive metrics, we used feature descriptors generated by the "molekulekit" utility. Through experimentation, we identified the optimal descriptors with a box size of 16 and voxel size of 0.5. This optimization elevated our results, leading to the creation of our model "AVGP3," which was subsequently integrated into our final ensemble.
Exploration of Rosetta scores: drawing inspiration from the notebook available at Deletion-specific ensemble, we observed the potential of Rosetta scores in predicting thermal stability. We integrated Rosetta scores with the "AVGP3" model in an ensemble configuration. Then we combined the "AVGP3" model with Deletion-specific ensemble using specific weight distributions (0.2 and 0.8).
Utilization of "Pose_Energies_Table": we meticulously examined the contribution of Rosetta scores by analyzing the "Pose_Energies_Table" in pdb files, which encapsulates 20 parameters and their associated weights. These parameters played a pivotal role in our approach.
Ranked Rosetta ensemble: An innovative approach emerged, which involved utilizing weights for the ensemble. Independent predictions of thermostability were generated for each parameter, individually, across all mutations. These individual predictions were then amalgamated into an ensemble configuration using the weights from the "Pose_Energies_Table." This ensemble, referred to as "RankedRosetta."
Final ensemble integration: a final result from community forums was integrated into our ensemble, boosting our results even more. The integration formula was expressed as (0.05 * Rasp + 0.95 * 0.615).
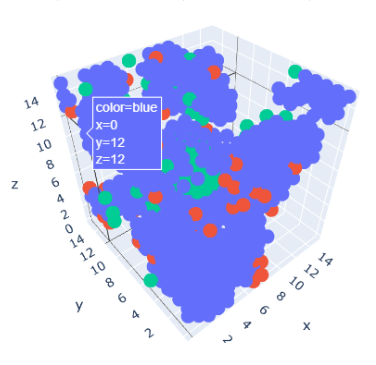
To summarize, the comprehensive approach of the QuData team has led to noticeable results. Our findings underscore the significant influence of Rosetta scores throughout the molecular structure, as well as the impact of the average hydrogen bond donor parameter in the vicinity of mutation sites on the prediction of thermostability.
